Error-correction DNA storage; nanopore storage.
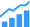
Error-correction DNA storage
Los Alamos National Laboratory has developed a key technology that could one day pave the way towards DNA storage.
Researchers have developed a technology called the Adaptive DNA Storage Codec (ADS Codex). ADS Codex is software that translates digital binary files into the four-letter genetic alphabet needed for DNA storage.
Deoxyribonucleic acid (DNA) is a molecule that carries the genetic instructions of living things. A complete set of DNA is called the genome.
DNA is also a promising archival storage technology. Several companies are working on the technology, which has been in R&D for years.
In computers, discrete units are stored as “0s” and “1s” as binary code. In contrast, DNA molecules encode information with sequences of discrete units. In DNA molecules, the units involve four distinct nucleotide bases: adenine (A), cytosine (C), guanine (G) and thymine (T).

Source: Los Alamos National Laboratory
DNA doesn’t degrade over time and is compact. So, it could be used to store a massive amount of data in a tiny space over a long period of time. For example, the Library of Congress has about 74 terabytes of data. Some 6,000 such libraries could fit in a DNA archive the size of a poppy seed, according to Los Alamos National Lab.
But DNA storage isn’t a mainstream technology. It’s expensive, slow and prone to mistakes. In response, Los Alamos is developing new technologies to overcome some of those issues. It’s part of a major program. Los Alamos’ work is part of the Intelligence Advanced Research Projects Activity (IARPA) Molecular Information Storage (MIST) program.
IARPA is an U.S. government group that provides R&D services for U.S. intelligence agencies. The goal of MIST is to write 1 terabyte—a trillion bytes—and read 10 terabytes within 24 hours for $1,000, according to Los Alamos.
That won’t be easy. Encoding a binary file into a molecule is done by DNA synthesis, according to Los Alamos. Then, ADS Codex translates the binary data into sequences of four letter combinations of A, C, G, and T. It also handles the decoding back into binary.
At times, though, DNA synthesis is prone to errors in the coding. To solve that issue, ADS Codex adds additional information called error detection codes. This in turn can be used to validate the data. Los Alamos has completed a version 1.0 of ADS Codex.
“On a digital hard disk, binary errors occur when a 0 flips to a 1, or vice versa, but with DNA, you have more problems that come from insertion and deletion errors,” said Latchesar Ionkov, a computer scientist at Los Alamos. “You’re writing A, C, G, and T, but sometimes you try to write A, and nothing appears, so the sequence of letters shifts to the left, or it types AAA. Normal error correction codes don’t work well with that.
“Appending a single nucleotide to DNA is very slow. It takes a minute. Imagine writing a file to a hard drive taking more than a decade. So that problem is solved by going massively parallel. You write tens of millions of molecules simultaneously to speed it up,” Ionkov said. “Our software, the Adaptive DNA Storage Codec (ADS Codex), translates data files from what a computer understands into what biology understands. It’s like translating from English to Chinese, only harder.”
Nanopore storage
Instead of DNA storage, the École polytechnique fédérale de Lausanne (EPFL) has developed a different biological approach to store data–bacterial nanopores.
EPFL has developed a nanopore-based system that can read data encoded into synthetic macromolecules with accuracy and resolution.
A nanopore is a tiny pore. Pores can be created in materials like silicon or biological entities like protein. In biotech, nanopores can be used for DNA sequencing or sensing biomolecules.
Nanopores, however, are limited in terms of resolutions. That presents some challenges if they are used as a storage medium.
In response, EPFL developed nanopores, based on a pore-forming toxin called aerolysin. Using aerolysin nanopores, researchers demonstrated the ability to decode binary information. They “adapted aerolysin to detect molecules tailored-made precisely to be read by this pore,” according to EPFL.
The molecules, known as digital polymers, were developed in the lab at the Institut Charles Sadron of the CNRS. “They are a combination of DNA nucleotides and non-biological monomers designed to pass through aerolysin nanopores and give out an electrical signal that could be read out as a bit,” according to EPFL.
To decode the readout signals, researchers used deep learning. This in turn allowed them to decode up to 4 bits of information from the polymers with high accuracy.
“But unlike conventional nanopore readouts, this signal delivered digital readings with single-bit resolution, and without compromising information density,” said Chan Cao, a researcher at EPFL.
“There are several improvements we are working on to transform this bio-inspired platform into an actual product for data storage and retrieval,” said Matteo Dal Peraro from EPFL. “But this work clearly shows that a biological nanopore can read hybrid DNA-polymer analytes. This opens up new promising perspectives for polymer-based memories, with important advantages for ultra-high density, long-term storage and device portability.”
Leave a Reply